LINKED PAPER
Near panmixia at the distribution-wide scale but evidence of genetic differentiation in a geographically isolated population of the Terek Sandpiper Xenus cinereus. Rönkä, N., Pakanen, V. M., Blomqvist, D., Degtyaryev, V., Golovatin, M., Isakov, G., Karlionova, N., Lehikoinen, A., Morozov, A., Paskhalny, S., Pauliny, A., Pinchuk, P., Rauhala, P., Tomkovich, P, Zakharov, E., Koivula, K. & Kvist, L.. 2019. IBIS. DOI: 10.1111/ibi.12651 VIEW
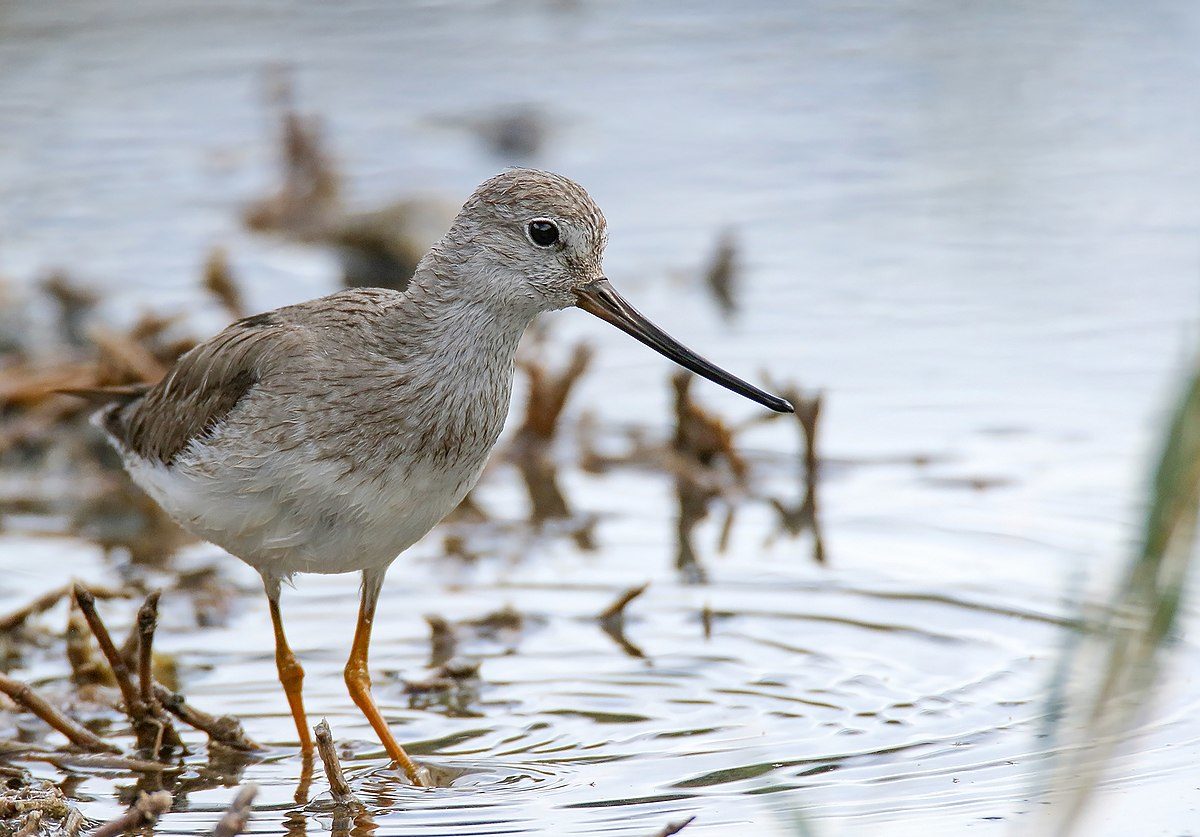
Genetic variation is not distributed randomly. When you compare the genetic make-up of several populations, you are bound to find some interesting geographical patterns. Isolation-by-distance, for example, indicates that the farther two populations are apart, the more they differ genetically. A related pattern is the so-called abundant-center hypothesis (Brussard 1984) which predicts that peripheral populations have lower genetic variation compared to populations in the center of the distribution. This pattern can arise due to a combination of inbreeding (peripheral populations are smaller) and fitness effects (conditions are better in the center compared to the edges). An international group of ornithologists tested this hypothesis for the Terek Sandpiper (Xenus cinereus), a small wader with a wide distribution.
Distribution
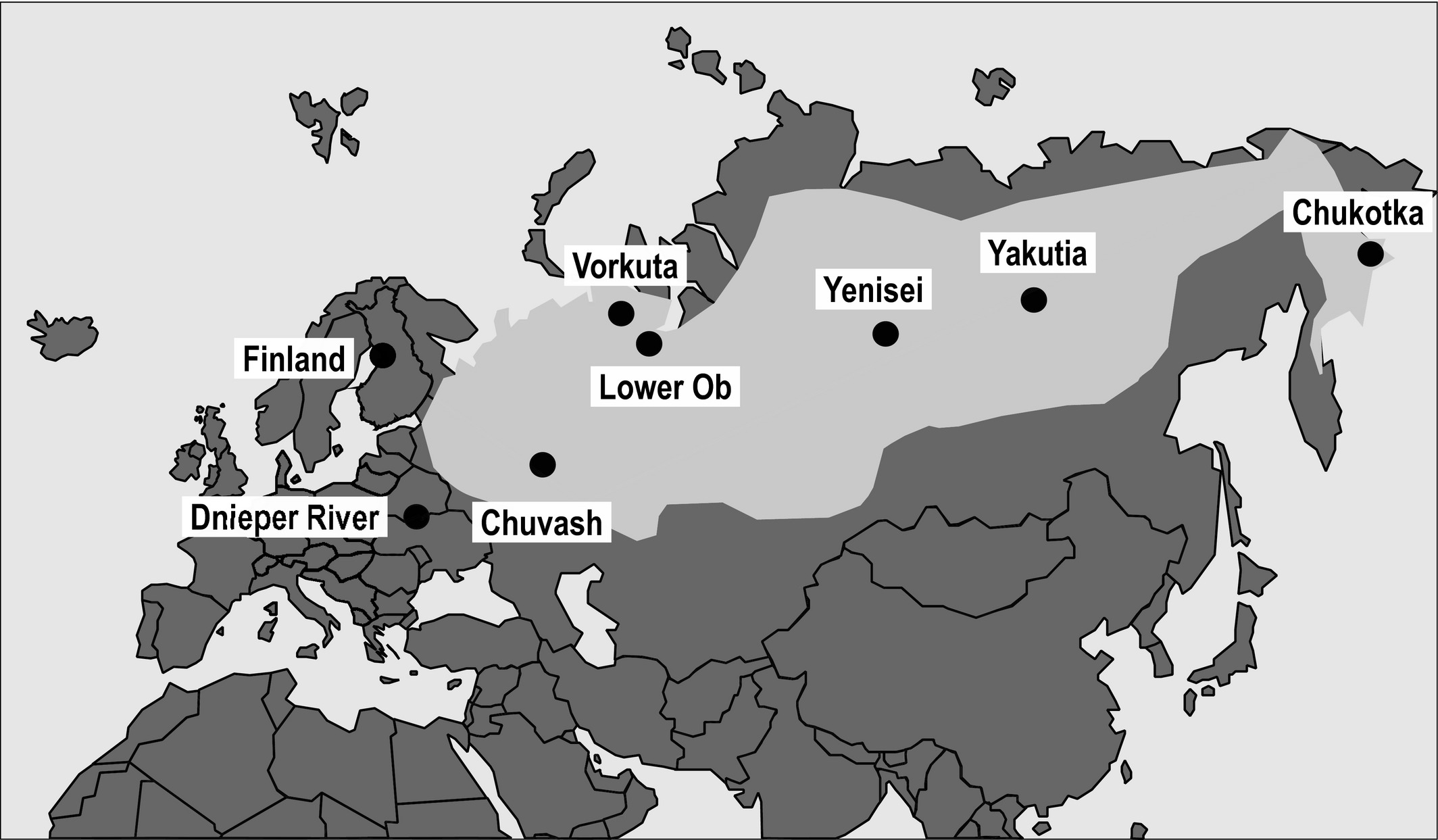
The Terek Sandpiper occurs from Finland to eastern Siberia. Although there are some differences in habitat use and wintering locations, this species is currently considered monotypic (i.e. no recognized subspecies). Its core breeding area ranges from the Volga River in the west to the large rivers of Siberia in the east. There are some peripheral populations in Finland and along the Dnieper River (Tomkovich et al. 2016). This geographic spread provides the ideal situation to test the abundant-center hypothesis outlined above. The researchers collected DNA samples from eight populations and studied genetic variation of 13 microsatellite loci and one mitochondrial marker.
Figure 1The main breeding distribution (light grey) and sampling sites (black circles) of the Terek Sandpiper.
Panmixia
The genetic analyses pointed to a lack of genetic population structure. Birds from different breeding areas might use the same wintering locations, facilitating genetic exchange between populations. Geneticists refer to this situation as panmictic (i.e. all individuals are potential partners). However, the absence of population structure could also reflect a recent population expansion after a severe genetic bottleneck (Bulgin et al. 2003). Perhaps the number of Terek Sandpipers was reduced during the last glaciation (ca. 10,000 years ago) and this species is currently recolonizing Eurasia. These scenarios will need to be explored with more fine-scale genomic data (Ottenburghs et al. 2019).
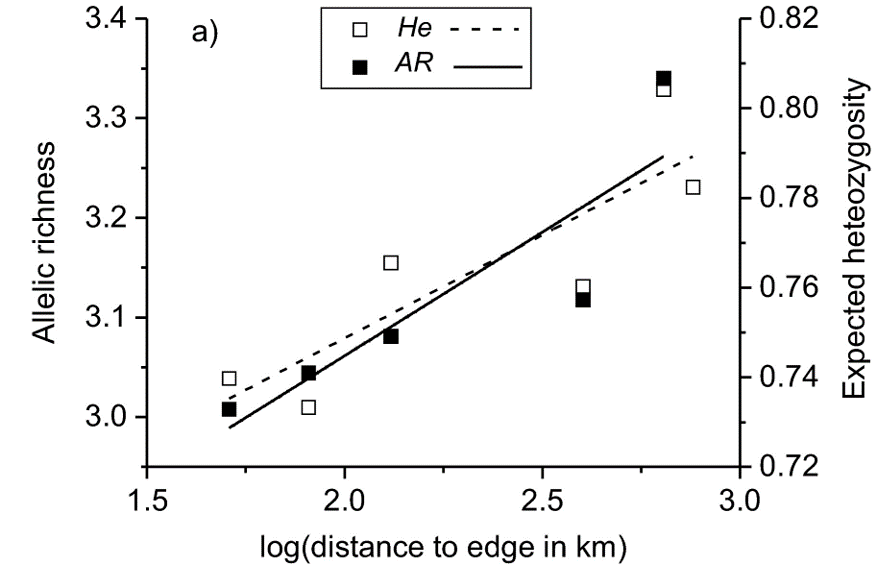
Figure 2 Relationship between the distance to the edge of the distribution (km, log‐transformed) and allelic richness (solid line) and expected heterozygosity (dotted line) of 13 microsatellite loci. The pattern conforms to the abundant-center hypothesis.
Genetic depletion
Despite the lack of population structure, the researchers did find evidence for the abundant-center hypothesis. Two measures of genetic diversity – allelic richness and expected heterozygosity – decreased towards the edge of the distribution. The lowest allelic richness was found in the isolated Dnieper River population, which was also the most differentiated population from the main distribution. This population was probably a recent addition to the already wide distribution of the Terek Sandpiper. The small population size and large distance from other populations render the Dnieper River population vulnerable to genetic depletion. Even though the Terek Sandpiper is considered of Least Concern by Birdlife International, it is important to monitor isolated populations and take conservation measures if needed (Moritz 1994).
References
Brussard, P. F. (1984). Geographic patterns and environmental gradients: the central-marginal model in Drosophila revisited. Annual Review of Ecology and Systematics 15: 25-64. VIEW
Bulgin, N. L., Gibbs, H. L., Vickery, P., & Baker, A. J. (2003). Ancestral polymorphisms in genetic markers obscure detection of evolutionarily distinct populations in the endangered Florida grasshopper sparrow (Ammodramus savannarum floridanus). Molecular Ecology 12: 831-844. VIEW
Moritz, C. (1994). Defining ‘evolutionarily significant units’ for conservation. Trends in Ecology & Evolution 9: 373-375. VIEW
Ottenburghs, J., Lavretsky, P., Peters, J. L., Kawakami, T., & Kraus, R. H. (2019). Population genomics and phylogeography. In Avian Genomics in Ecology and Evolution (pp. 237-265). Springer, Cham. VIEW
Tomkovich, P.S., Svirodova, T.V., Kossenko, S.M., Mischenko, A.L. & Nikolaev, V.I. (2016). Does the Terek Sandpiper Xenus cinereus have an unbroken breeding range in western Russia? Ornithologia 40: 101-109. VIEW
Image credits
Featured image: Terek Sandpiper Xenus cinereus | Imran Shah | CC BY-SA 2.0 Wikimedia Commons